As a reminder the first post concerns doing a merging scheme using the lapply function. After that in the second post we looked how it translates to a parallel solution for even more speed.
As a reminder the problem is the following: given N amount of files with a random amount of variables in each file. How do we merge these files into a complete data within reasonable amount of time? For example you can consider that the medical instruments records it's each run (patient) to a different file. With only the values measured by the instrument are recorded to the file.
In practice I'm using Eve Online killmails. So far I have around 160k mails. Hence the need for a fast solution.
Personally I haven't really used data.table, but it looks good enough. So I recommend checking it out. But in a sense nothing's really changed on my code part. I did a fast merging function based on data.table merge called the lady, but won't give the code because it isn't fast enough (spoiler alert). Also the data.table merge is a generic function. Meaning if you are using data.tables in a merge it automaticaly uses the data.table merge function. But enough it this prattle. Here's the results.
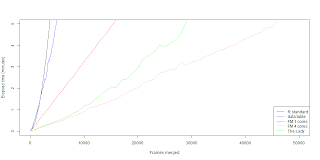
data.table merge is faster then the R core merge function if you are merging a for-loop solution. The problem still is that the data.table merging in a for-loop works in exponential time. Which doesn't translate well for big data. The single core fast merging scheme still beats the data.table merge in a for-loop. The biggest wonder is why when we combine the data.table merge with parallel fast merging scheme we get a slower outcome. Naturally I though we could squeeze few seconds out with low frame count, but seems that it's actually few minutes worse. So far I'm running dry in practical ideas and reasons why the lady seems so fat. Maybe it doesn't cleane up it's diet ? (memory management?) If you have some insight please comment.
As requested, here's the bench.
How are you doing the merge with ```data.table```? I've found that doing ```A[B]``` is much faster than ```merge(A, B)```.
ReplyDeleteIt is quite stressful to see a chart like that. Can you please post a single reproducible block of code that you used to create it? Then we can investigate. If it is wrong then I hope you will publish a correction.
ReplyDeleteMatthew Dowle (main author of data.table)
Hi Dowle,
DeleteI hope my newest post is to your satisfaction. It is not the one I made the plot, but you can use it to replicate my point (hopefully).
Hmm.... I might as well add the mark3 script to this data.table vs core merge post.
- Xachriel
Hi,
ReplyDeleteThanks, getting there. But it fails with the error :
object 'eve.data.frame' not found
A reproducible example means something someone else can paste into a fresh R session and it produces that plot. For example, the types of the columns in eve.data.frame could be the problem (if they are character for example). But I can't check that if you don't provide it. When you post reproducible code to the internet it's good practice to first paste the code into a fresh R session on your side, as a final check.
Btw, subscribing to updates on your blog doesn't seem to work. So I've set a reminder to come back here and check. For a faster response please email me directly or ask on datatable-help or Stack Overflow.
Matthew Dowle
I tried again to subscribe. I'm logged in to your site as myself. I click on "Subscribe to: Post Comments (Atom)" at the bottom. The screen flashes but nothing else happens as far as I can see. I'm using Google Chrome 21.0.1180.89m.
ReplyDeleteMatthew
hmm... I did spawn a new session through notepad++ and it works correctly. Did you unpackage the data and the working directory correctly?
DeleteAlso this is just the end part for the bench, you have to define the data using the test bench. I'll add the data defining parts here too.
The defining data step is 50 lines. Felt it added unnecessary length. But, here you go.
I'll do something about that subscription thing. I'm fairly new, busy and lazy. The killer combination... I'm less busy tomorrow, so I'll lookit up then.
Did a completely new blogger account and subscription worked... Still error ?
DeleteI think this is an informative post and it is very useful and knowledgeable. therefore, I would like to thank you for the efforts you have made in writing this article. big data analytics
ReplyDeletethanks this is good blog. privacyenbescherming
ReplyDelete